BESA Research 7.1 January 2024 is now released. We’re thrilled to announce this latest release of our software, packed with enhancements to elevate your experience. Please make sure that you update to this version! Particular highlights are:
General:
Windows 11 Support: Embrace the latest technology with full compatibility for Windows 11. Please note, support for Windows 7 has been deprecated in this version.
Data Review and Pre-processing:
- Event Histogram: Visualize trigger event frequency (e.g. of epileptic spikes) in long EEG recordings effortlessly in histograms: Use menu View / Histogram or type the key H on the keyboard. In case that tags were marked for spike events, first run the batch ‘Epilepsy/F6_Spike_Convert_Tags_And_Average’ to create triggers from the tags (or press F6 after setting BESA Research to EEG Review mode using Options / Reset Settings)
- Pattern Search: When searching with Query, visualize each candidate event as a 3D map to streamline your search process
- Expanded Data Formats: We now support additional file formats:
- OPM by Cerca Magnetics (using a proprietary script cMEG2fif.py to transform data to fif): Note that this option requires the purchase of an additional module license. Signal subspace projection can be applied to reduce external noise. The full analysis functionality is available.
- MED Dark Horse
- Data reader improvements: BESA Research can now read Micromed version 5, XLTEK version 9.3, g.Tec HDF5, and Curry 9
- Improved Export: Experience enhanced data export to EDF format and improved epoch handling around triggers
- Batch Processing:
- Customizable Defaults: Set default values for user-defined batch variables
- Expanded Functionality:
- DisplayMRI now allows showing the cortex viewx
- SetInterval can automatically set a spike onset interval
- SaveBitmap now enables saving as .bmp or .png format
- SchmittTrigger auto-generates triggers from EMG data or from a trigger channel
- Enhanced Compatibility: Our software now supports the latest MATLAB versions seamlessly
- ERP Processing: Define more complex conditions with an increased boolean expression length for conditions
- ini File customization options: Tailor your experience with addional parameters, including OPM data default scaling, and enhanced filter control options
MEG Version:
Enhanced Mapping: Experience improved FFT mapping for MEG with support for 3D mapping
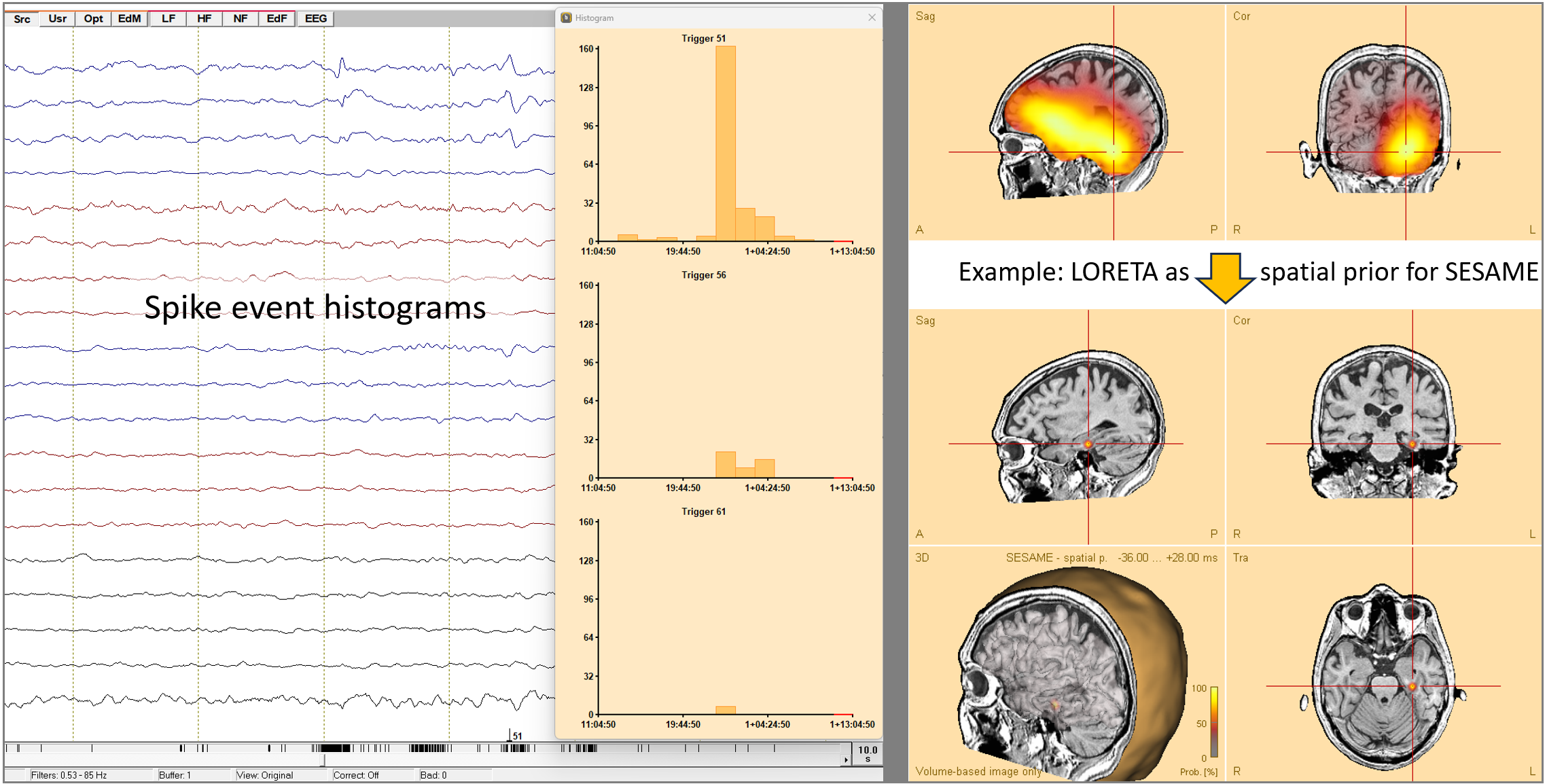
Source Analysis and Source Imaging:
- SESAME:
- Utilize spatial priors effectively with SESAME, enhancing source imaging accuracy. To use them: Compute or load any source image in the Source Analysis window. Then press the button ‘Weight by Image’ before you start SESAME computation
- The hyperpriors integration was improved. Default settings were changed to run SESAME for a fixed 50 iterations since this enables convergence in nearly all cases
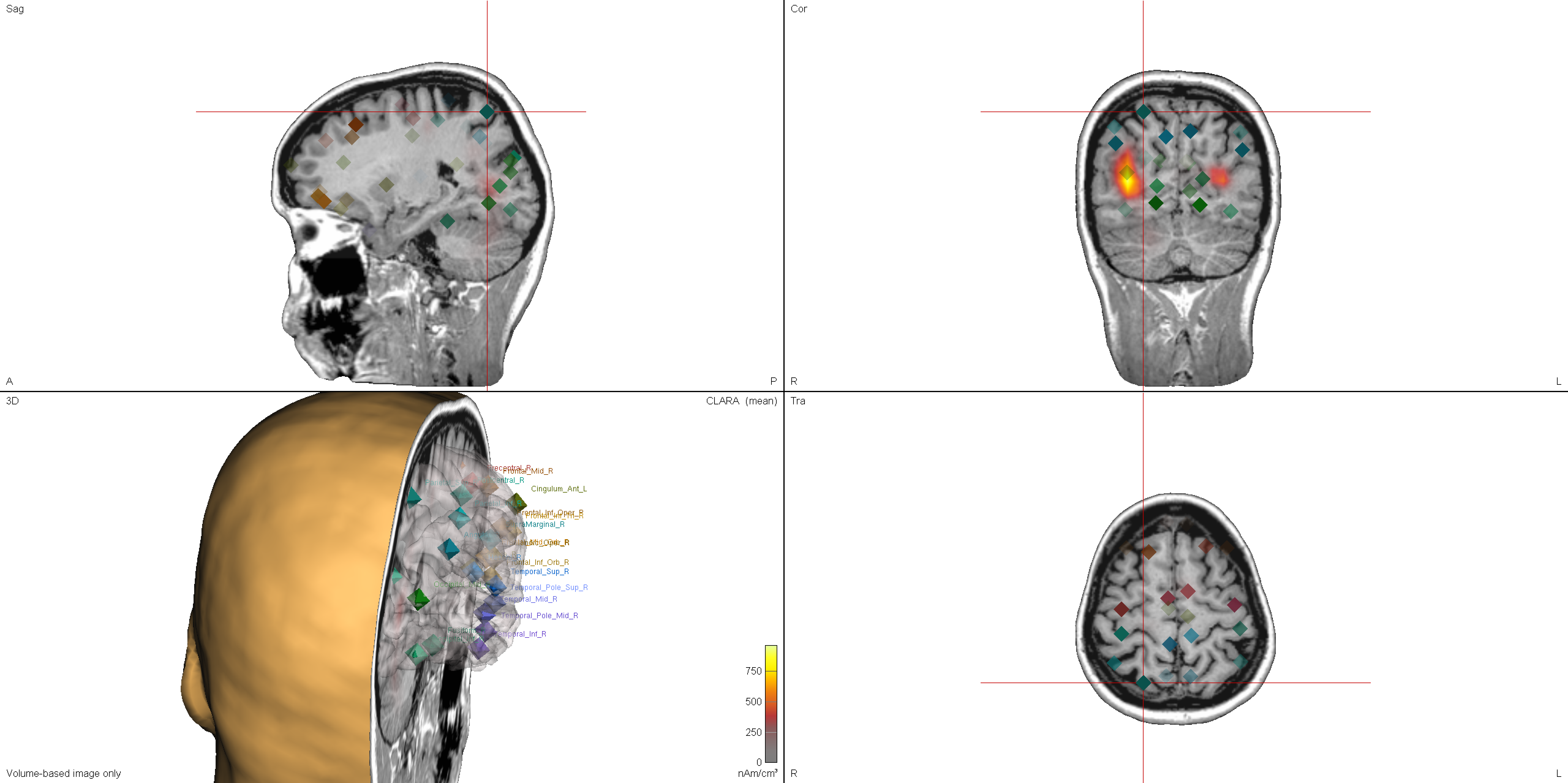
- 3D window: Source labels can now be displayed beside the source symbols. Use the context menu to toggle the display (default: off)
- Export Options: Export source images effortlessly to high-resolution AC-PC NIfTI files, ensuring seamless integration with individual anatomy (also available as batch export)
- Solution Integrity: Baseline information is now preserved for solutions saved in specific coordinate systems, ensuring correct re-computation of statistical parameters when re-loading solutions
- Screenshots of the source analysis and / or 3D window can now be saved as .png or .bmp files, with increased resolution
Source Coherence window:
- Optimization: Source Coherence module now leverages OpenGL acceleration by default, ensuring faster performance
- Large-data computation: Data buffers for time-frequency computations were optimized to allow for larger datasets to be stored in memory
- User Interface: The Source Coherence window now stays visible after computing the time-frequency beamformer, facilitating parameter adjusting before re-computing
Besides, this release contains many bugfixes. For a full list of all changes, see here.

Comments are closed.